Overview
Whales and dolphins make up the cetaceans, a special group of mammals adapted to life underwater. Their transition from dry land to an aquatic environment is one of the greatest examples of evolutionary adaptation. Georgia Tsagkogeorga and her colleagues at Queen Mary University of London were interested in how cetaceans evolved in comparison with their closest relatives. In particular, they wanted to know if the molecular adaptations related to aquatic lifestyle appeared before or after whales and dolphins split from the hippos. Using GridPP resources, the team were able to test different evolutionary hypotheses on a vast scale to gain an insight.
A humpback whale, one of the species featured in the study.
The problem
To test whether selection pressures acted on a given gene, a series of probabilistic analyses using codon models of evolution need to be performed. This involves fitting two models onto the data: one representing the null neutral evolution hypothesis and a second model assuming selective pressures acting on the gene. By comparing the likelihoods of the null vs. alternative models, the selective regime of the gene in the lineage of interest can be inferred. However, this needed to be done for 11,000 genes across five lineages of aquatic and semi-aquatic mammals.
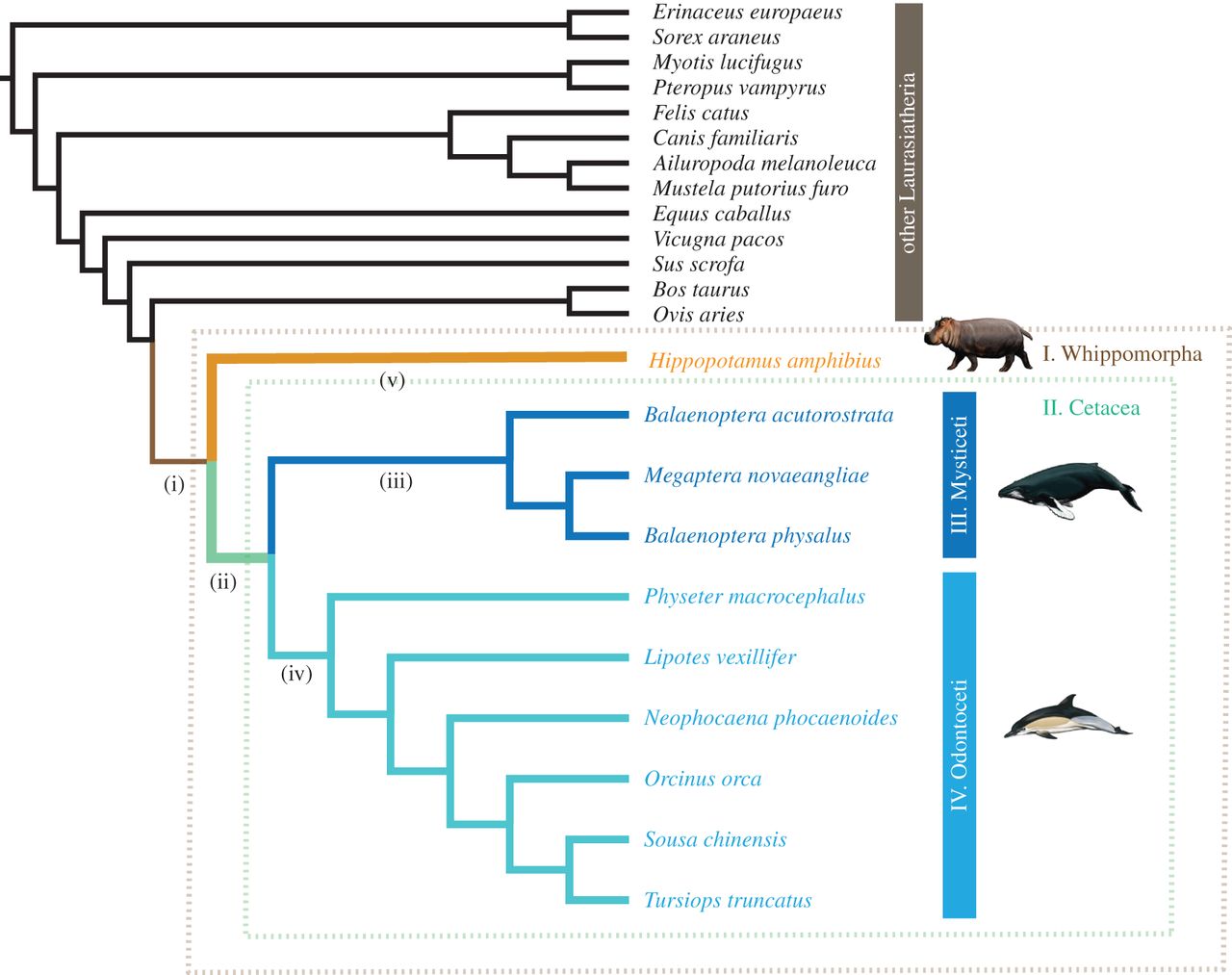
Evolutionary relationships among laurasiatherian mammals as used in molecular evolution analyses performed with GridPP resources. Figure reproduced from (Tsagkogeorga, 2015).
The solution
Fortunately the required analyses were highly parallelisable, making them ideal for running on the Grid. With support from the Queen Mary University of London Tier-2 cluster team, Tsagkogeorga and her colleagues were able to run the required 110,000 computations quickly and efficiently. The conclusions, published in Royal Society Open Science, show that the most significant molecular adaptations to aquatic life, for example the evolution of underwater sensory perception or their ability to dive, appeared in cetaceans after the split with the hippos, 55 million years ago.
What they said
These analyses would have taken months to complete without access to parallel computing. Our study benefited greatly from the 3,000+ CPU cores of QMUL GridPP cluster, by speeding up the analyses to only a few weeks or even days.Dr Georgia Tsagkogeorga, School of Biological and Chemical Science, QMUL
Further details
Services used
- Compute: The studies were run on the High Throughput Computing (HTC) GridPP cluster at QMUL;
- Storage: Results were stored on the QMUL cluster’s Lustre-based storage system.
Supporting GridPP sites
- Queen Mary University of London
Publications
- G Tsagkogeorga, “A phylogenomic analysis of the role and timing of molecular adaptation in the aquatic transition of cetartiodactyl mammals“, R. Soc. open sci. 2 150156 (2015)
Acknowledgements
In addition to the support provided by STFC for GridPP, this work was funded by the European Research Council (ERC 1076 Starting grant no. 310482). This case study originally featured on the EGI website.